Community Articles
- Cloudera Community
- Support
- Community Articles
- NiFi in Healthcare - Ingesting HL7 Data in NiFi
- Subscribe to RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
- Subscribe to RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
Created on
09-22-2017
09:03 PM
- edited on
02-17-2020
10:21 AM
by
VidyaSargur
Ingesting HL7 Data in NiFi
Pre-requisites
We will assume you have the following:
- Basic understanding of Linux CLI
- HDF cluster or sandbox with NiFi, Kafka, and HDFS installed and running
- Basic knowledge of connecting components in NiFi and configuring settings
Overview
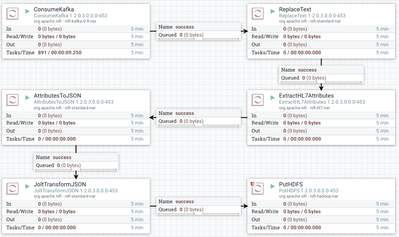
Here's the finished product:
This NiFi flow accomplishes the following:
- Read the data from a Kafka stream
- Format the data to comply with HL7 specifications
- Extract the HL7 attributes from the formatted content block
- Transform the attribute list into a JSON block
- Impose the data's natural hierarchy onto the JSON rather than leaving it as a flat-mapped structure
- Write the data to HDFS
Here's an example of our input data:
MSH|^~\&|XXXXXX||HealthOrg01||||ORU^R01|Q1111111111111111111|P|2.3|<cr>PID|||000000001||SMITH^JOHN||19700101|M||||||||||999999999999|123456789|<cr>PD1||||1234567890^LAST^FIRST^M^^^^^NPI|<cr>OBR|1|341856649^HNAM_ORDERID|000000000000000000|648088^Basic Metabolic Panel|||20150101000100|||||||||1620^Johnson^John^R||||||20150101000100|||M|||||||||||20150101000100|<cr>OBX|1|NM|GLU^Glucose Lvl|159|mg/dL|65-99^65^99|H|||F|||20150101000100|
And the output data after being written to HDFS:
{
"OBX_1": {
"UserDefinedAccessChecks": "20150101000100",
"ObservationIdentifier": {
"Text": "Glucose Lvl",
"Identifier": "GLU"
},
"ReferencesRange": "H",
"Units": {
"NameOfCodingSystem": "99",
"Identifier": "65-99",
"Text": "65"
},
"ObservationSubID": "159",
"NatureOfAbnormalTest": "F",
"SetIDOBX": "1",
"ValueType": "NM",
"ObservationValue": "mg\/dL"
},
"OBR_1": {
"OrderingProvider": {
"FamilyName": "Johnson",
"IDNumber": "1620",
"GivenName": "John",
"MiddleInitialOrName": "R"
},
"UniversalServiceIdentifier": {
"Text": "Basic Metabolic Panel",
"Identifier": "648088"
},
"FillerOrderNumber": {
"EntityIdentifier": "000000000000000000"
},
"PlacerOrderNumber": {
"NamespaceID": "HNAM_ORDERID",
"EntityIdentifier": "341856649"
},
"ResultStatus": "M",
"ObservationDateTime": "20150101000100",
"ScheduledDateTime": "20150101000100",
"SetIDObservationRequest": "1",
"ResultsRptStatusChngDateTime": "20150101000100"
},
"MSH": {
"MessageControlID": "Q1111111111111111111",
"SendingApplication": {
"NamespaceID": "XXXXXX"
},
"ReceivingApplication": {
"NamespaceID": "HealthOrg01"
},
"ProcessingID": {
"ProcessingID": "P"
},
"MessageType": {
"MessageType": "ORU",
"TriggerEvent": "R01"
},
"EncodingCharacters": "^~\&",
"VersionID": "2.3",
"FieldSeparator": "|"
},
"uuid": "de394ca2-cbaf-4703-9e25-4ea280a8c691",
"PID": {
"SSNNumberPatient": "123456789",
"PatientAccountNumber": {
"ID": "999999999999"
},
"DateOfBirth": "19700101",
"Sex": "M",
"PatientName": {
"GivenName": "JOHN",
"FamilyName": "SMITH"
},
"PatientIDInternalID": {
"ID": "000000001"
}
},
"path": ".\/",
"PD1": {
"PatientPrimaryCareProviderNameIDNo": {
"IDNumber": "1234567890",
"FamilyName": "LAST",
"GivenName": "FIRST",
"AssigningAuthority": "NPI",
"MiddleInitialOrName": "M"
}
},
"filename": "279877223850444",
"kafka": {
"partition": "0",
"offset": "221",
"topic": "test"
}
}
Reading from Kafka
To read from Kafka, we will need to first create a topic. Execute the following to create a topic:
./kafka-topics.sh --create --zookeeper <ZOOKEEPER_HOSTNAME>:2181 --replication-factor 1 --partitions 1 --topic test
You should have a topic named test available to you.
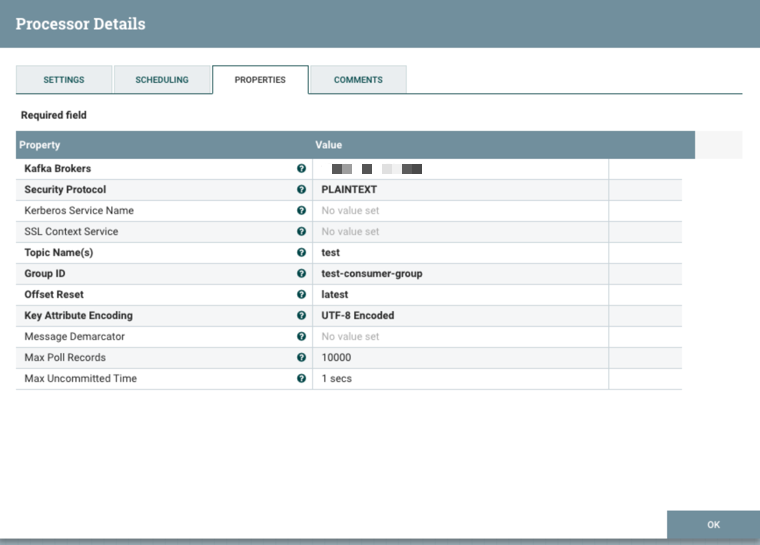
We can now create the ConsumeKafka processor with this configuration:
You can test that this topic is functioning correctly by using the command-line kafka-console-producer and kafka-console-consumer tools in two different command prompts.
Formatting to Spec
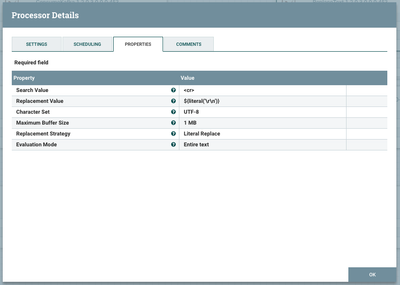
The text data we are using had to be compressed into one line since we are using Kafka to read it (Kafka needs each entry as a single line without any carriage returns), so we now need to de-compress our FlowFile into the appropriate number of lines. We will be replacing all '<cr>' strings with a carriage return ('\r', 0x0D). Use the following configuration on the ReplaceText processor to do so:
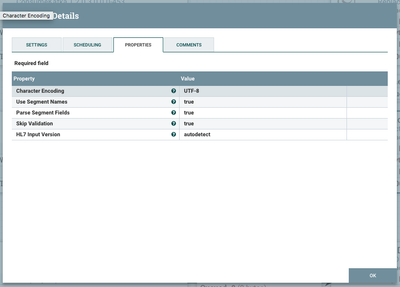
Extracting HL7 attributes
We will use the built-in ExtractHL7Attributes processor to transform our formatted text into "Attributes" that are native to NiFi and understood by it.
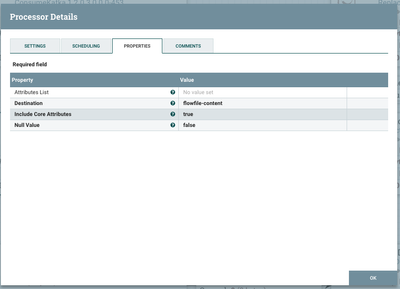
Transforming into JSON
We can now do a simple conversion of those attributes into JSON. Note that our data is destined for the flowfile-content, so we will now overwrite the formatted data that we used to extract our attributes from. Up until now, we had both the content and attributes which are duplicates of the content.
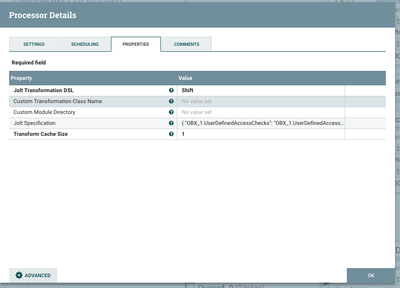
Formatting the JSON
To format the JSON, we will use a Jolt Shift specification. This specification will fully depend on the data you are using and expecting. The sample spec below is based on my sample input data.
{
"OBX_1.UserDefinedAccessChecks": "OBX_1.UserDefinedAccessChecks",
"OBR_1.OrderingProvider.FamilyName": "OBR_1.OrderingProvider.FamilyName",
"MSH.MessageControlID": "MSH.MessageControlID",
"OBX_1.ObservationIdentifier.Text": "OBX_1.ObservationIdentifier.Text",
"MSH.SendingApplication.NamespaceID": "MSH.SendingApplication.NamespaceID",
"OBR_1.UniversalServiceIdentifier.Text": "OBR_1.UniversalServiceIdentifier.Text",
"MSH.ReceivingApplication.NamespaceID": "MSH.ReceivingApplication.NamespaceID",
"MSH.ProcessingID.ProcessingID": "MSH.ProcessingID.ProcessingID",
"uuid": "uuid",
"PID.SSNNumberPatient": "PID.SSNNumberPatient",
"OBR_1.FillerOrderNumber.EntityIdentifier": "OBR_1.FillerOrderNumber.EntityIdentifier",
"path": "path",
"PID.PatientAccountNumber.ID": "PID.PatientAccountNumber.ID",
"PID.DateOfBirth": "PID.DateOfBirth",
"PD1.PatientPrimaryCareProviderNameIDNo.IDNumber": "PD1.PatientPrimaryCareProviderNameIDNo.IDNumber",
"PID.Sex": "PID.Sex",
"MSH.MessageType.MessageType": "MSH.MessageType.MessageType",
"OBX_1.ReferencesRange": "OBX_1.ReferencesRange",
"OBR_1.OrderingProvider.IDNumber": "OBR_1.OrderingProvider.IDNumber",
"PD1.PatientPrimaryCareProviderNameIDNo.FamilyName": "PD1.PatientPrimaryCareProviderNameIDNo.FamilyName",
"OBX_1.Units.NameOfCodingSystem": "OBX_1.Units.NameOfCodingSystem",
"OBX_1.Units.Identifier": "OBX_1.Units.Identifier",
"filename": "filename",
"PID.PatientName.GivenName": "PID.PatientName.GivenName",
"OBX_1.ObservationSubID": "OBX_1.ObservationSubID",
"PD1.PatientPrimaryCareProviderNameIDNo.GivenName": "PD1.PatientPrimaryCareProviderNameIDNo.GivenName",
"OBR_1.PlacerOrderNumber.NamespaceID": "OBR_1.PlacerOrderNumber.NamespaceID",
"MSH.MessageType.TriggerEvent": "MSH.MessageType.TriggerEvent",
"PD1.PatientPrimaryCareProviderNameIDNo.AssigningAuthority": "PD1.PatientPrimaryCareProviderNameIDNo.AssigningAuthority",
"OBR_1.ResultStatus": "OBR_1.ResultStatus",
"PID.PatientName.FamilyName": "PID.PatientName.FamilyName",
"MSH.EncodingCharacters": "MSH.EncodingCharacters",
"MSH.VersionID": "MSH.VersionID",
"kafka.partition": "kafka.partition",
"OBR_1.UniversalServiceIdentifier.Identifier": "OBR_1.UniversalServiceIdentifier.Identifier",
"OBR_1.ObservationDateTime": "OBR_1.ObservationDateTime",
"OBR_1.ScheduledDateTime": "OBR_1.ScheduledDateTime",
"OBX_1.ObservationIdentifier.Identifier": "OBX_1.ObservationIdentifier.Identifier",
"OBR_1.OrderingProvider.GivenName": "OBR_1.OrderingProvider.GivenName",
"OBR_1.SetIDObservationRequest": "OBR_1.SetIDObservationRequest",
"OBR_1.ResultsRptStatusChngDateTime": "OBR_1.ResultsRptStatusChngDateTime",
"OBR_1.PlacerOrderNumber.EntityIdentifier": "OBR_1.PlacerOrderNumber.EntityIdentifier",
"OBX_1.NatureOfAbnormalTest": "OBX_1.NatureOfAbnormalTest",
"OBX_1.SetIDOBX": "OBX_1.SetIDOBX",
"MSH.FieldSeparator": "MSH.FieldSeparator",
"PD1.PatientPrimaryCareProviderNameIDNo.MiddleInitialOrName": "PD1.PatientPrimaryCareProviderNameIDNo.MiddleInitialOrName",
"OBX_1.Units.Text": "OBX_1.Units.Text",
"OBX_1.ValueType": "OBX_1.ValueType",
"kafka.offset": "kafka.offset",
"PID.PatientIDInternalID.ID": "PID.PatientIDInternalID.ID",
"kafka.topic": "kafka.topic",
"OBX_1.ObservationValue": "OBX_1.ObservationValue",
"OBR_1.OrderingProvider.MiddleInitialOrName": "OBR_1.OrderingProvider.MiddleInitialOrName"
}
Here's the associated NiFi configuration:
Writing to HDFS
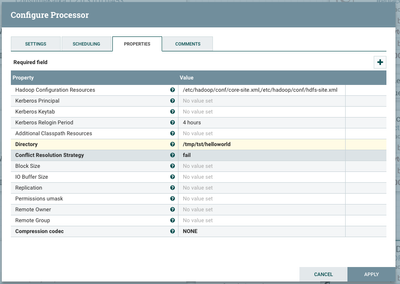
Finally, we will persist this JSON-transformed data to HDFS for further analysis and storage. Configure your PutHDFS processor as follows
Note that the 'Hadoop Configuration Resources' is a required field for your connection from NiFi to HDFS to work properly, so be sure to fill that in with wherever your core-site.xml and hdfs-site.xml HDFS configuration files are on disk.
Be sure to create the directory /tmp/tst/helloworld in hdfs and change ownership to the nifi user:
hdfs dfs -mkdir -p /tmp/tst/helloworld hdfs dfs -chown -R nifi /tmp/tst
Putting it all together
Now you should be able to connect all of your processors, start them, and see your data move through.
To send the sample data through our new flow, do the following:
1. SSH into a kafka broker in your cluster and cd to the binaries folder - in my environment, that is located at /usr/hdp/current/kafka-broker/bin.
2. Create a file in your home directory and paste the contents of your sample input data (from the Overview section above). This file should be exactly 1 line, no more.
3. Feed your sample data into the kafka producer with the following command:
/usr/hdp/current/kafka-broker/bin/kafka-console-producer.sh --broker-list <KAFKA_BROKER_HOSTNAME>:6667 --topic test < hl7data.txt
If all goes well, when you NiFi flow refreshes you should see it go all the way through. At this point you can check HDFS with the following command:
hdfs dfs -ls /tmp/tst/helloworld
You should see a file there. You can read it with the following:
hdfs dfs -cat /tmp/tst/helloworld/<FILENAME>
Thanks for reading!
Full NiFi template: hl7flow-template.xml
Created on 09-21-2018 02:53 PM
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
As I am new to Kafka I followed the steps for installing Kafka. Also tested Kafka using the Kafka Producer and Consumer bat file.
As a next step instead of Kafka Consumer console, I used NiFi ConsumeKafka Processor as a Consumer. Whatever data I send from Kafka Producer console I am able to receive it in ConsumeKafka Processor which I tested by sending the data through PutFile Processor. So as per my understanding the Topic name mentioned in the Producer console application and the ConsumeKafka Processor matches the topic name. Hence the Processor is able to read the message from the stream.
In my scenario I want to receive the HL7 message from a HL7simulator or through some asset which send the HL7 message to the configured IP:Port. I tried receiving HL7 message through ListenTCP or GetTCP with both of these Processor. But I am unable to receive the HL7 message in NiFi.
So I tried to follow the steps mentioned here (I thought If I use consumeKafka I may able to receive the HL7 message). But here the problem is I don't have a configuration section where I can mention topic. I can only set the IP, Port and Message in the simulator app.
Any help on receiving HL7 message into NiFi will be really helpful.
Thanks..
Created on 10-02-2018 11:19 PM
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Nice article - very well illustrated and explained. Wanted to try this out without Kafka and HDF and locally within NiFi. So modified the flow with GenerateFlowFile and PutFile processors. Worked great! Attached is the template - .hl7-processingexample.xml
Created on 10-03-2018 12:26 AM
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
@Dhamotharan P Why did the PutTCP and ListenTCP not work. Please see the attached template for working the same example using TCP connections. tcp-hl7-example.xml
Created on 10-03-2018 02:07 PM
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
@Muki Soomar Thanks for the template it will help me during my implementation. But I am stuck with the first step that is listening for the message over ListenTCP Processor. In the template which you shared It has generate flow file which send HL7 message and it flow through PutTCP. This work for me. But instead of generate flow file I tried using different HL7 simulators. But I am not able to receive in ListenTCP Processor. Now I am in the process of creating a custom processor on top of ListenTCP which will receive the HL7 message and also send back the acknowledgement.
Created on 10-29-2018 10:25 PM
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Because HL7 uses MLLP which is a protocol above TCP. I wonder if there is a plan to add MLLP support to NiFi.
Created on 11-13-2018 06:02 PM
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
@Dhamotharan P Could you let me know which simulator you were using so I can take a look at it?
Created on 11-15-2018 07:54 AM
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Hi @Muki Soomar Thanks for extending your help on this. I have created the custom processor to solve the above issue. But the answer for your question and the issue which I currently stuck with during HL7 message processing are posted in the link https://community.hortonworks.com/questions/226511/extracthl7attribute-processor-throws-error-cauhnh.... Please have a look on that and let me know your inputs on that. Thanks again..
Created on 06-11-2021 05:02 AM
- Mark as Read
- Mark as New
- Bookmark
- Permalink
- Report Inappropriate Content
Please how can i make sure that data on the edge device are anonymized and the identity of the patients protected?