Community Articles
- Cloudera Community
- Support
- Community Articles
- Part 3: Use Spark and Zeppelin to train fatigue pr...
- Subscribe to RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
- Subscribe to RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
Created on
01-31-2019
05:13 PM
- edited on
04-21-2026
04:38 AM
by
GrazittiAPI
Introduction
It's been a while, but I'm finally finishing this series on Best Mode Quotient. This article is fairly straight forward as we look how to build the infrastructure of model training using Zeppelin and Spark.
Pre-Requisites
In order to run this install, you will have to have deployed a BMQ ephemeral cluster as detailed in this article and this repository.
Moreover you will have to have gathered data on your fitness as explained in part 1 of these series. Based on this data, I created 3 indexes:
- Intensity Index: How intense was the workout I did that day (based on distance, pace and elevation)
- Fatigue Index: How much rest I had on that day (based on hours of sleep and rest heart rate)
- BMQ Index: How my BMQ fairs compare to the max BMQ (5)
You can then agglomerate these 3 indexes, having BMQ and Fatigue on the same day correlating to the Intensity Index of the previous day. All data will be shared in the last article of the series.
You will also notice that the model uses parameterized sleep and rest HR. The whole flow is to be revealed in part 4 🙂
Agenda
This tutorial is divided in the following sections:
- Section 1: Create a mysql interpreter for JDBC in Zeppelin
- Section 2: Create training set for BMQ prediction
- Section 3: Create a prediction model
- Section 4: Save the results in a table
Section 1: Create a mysql interpreter for JDBC in Zeppelin

Login to Zeppelin, then go to top right corner > Admin > Interpreter and edit the jdbc interpreter.
Add the following parameters:
- mysql.driver:
com.mysql.jdbc.Driver - mysql.url:
jdbc:mysql://localhost:3306/beast_mode_db - mysql.user:
bmq_user - mysql.password:
Be@stM0de - Add artifact:
mysql:mysql-connector-java:5.1.38
Restart the interpreter and you should be good to go.
Section 2: Create training set for BMQ prediction
Create a new note called BMQ Predictions and add the following code, using the jdbc interpreter you just built.
Delete existing training tables if any
%jdbc(mysql) drop table if exists training_set
Create training set based on fatigue and intensity indexes
%jdbc(mysql) create table training_set as ( select @rowid:=@rowid+1 as rowid, bmq_index.date, bmq_index, fatigue_index, intensity_index from bmq_index, fatigue_index, intensity_index, (select @rowid:=0) as init where bmq_index.date = fatigue_index.date and date_sub(bmq_index.date, INTERVAL 1 DAY) = intensity_index.date order by bmq_index.date asc)
View Data
%jdbc(mysql) select * from training_set
Delete existing prediction tables if any
%jdbc(mysql) drop table if exists prediction
Create a table we want to apply the algo against
%jdbc(mysql) create table prediction as ( select date(training_date) as date, estimated_intensity_index, round(( (1-((select (sleep_hours*60) from PREDICTION_PARAMETERS)/(select max(TOTAL_MINUTES_ASLEEP) from SLEEP_HISTORY)))*0.6 + (1-((select min(REST_HR) from HEALTH_HISTORY)/(select rest_hr from PREDICTION_PARAMETERS)))*0.4 ) *100,2) as estimated_fatigue_index, 0.0 as predicted_bmq from training_plan)
View Data
%jdbc(mysql) select * from prediction
Section 3: Create a prediction model
DISCLAIMER: This model needs to be worked on; the purpose of this article is to establish the principal architecture, not give the final most tuned model as I plan on improving on it.
This part uses the spark interpreter of Zeppelin to vectorize, normalize and train a model
Create dataframe from MySQL tables:
%spark2
import org.apache.spark.sql.SQLContext
import org.apache.spark.ml.regression.LinearRegression
import org.apache.spark.ml.feature.Interaction
import org.apache.spark.ml.feature.VectorAssembler
import org.apache.spark.ml.feature.Normalizer
val sqlcontext = new org.apache.spark.sql.SQLContext(sc)
val df = sqlcontext.read.format("jdbc").option("url", "jdbc:mysql://localhost:3306/beast_mode_db").option("driver", "com.mysql.jdbc.Driver").option("dbtable", "training_set").option("user", "bmq_user").option("password", "Be@stM0de").load()
df.show()
%spark2
val target_df = sqlcontext.read.format("jdbc").option("url", "jdbc:mysql://localhost:3306/beast_mode_db").option("driver", "com.mysql.jdbc.Driver").option("dbtable", "prediction").option("user", "bmq_user").option("password", "Be@stM0de").load()
target_df.show()Vectorize Dataframes
val assembler1 = new VectorAssembler().
setInputCols(Array( "fatigue_index","intensity_index")).
setOutputCol("features").
transform(df)
assembler1.show()%spark2
val assembler2 = new VectorAssembler().
setInputCols(Array( "estimated_fatigue_index","estimated_intensity_index")).
setOutputCol("features").
transform(target_df)
assembler2.show()Normalize Dataframes
%spark2
val normalizer = new Normalizer()
.setInputCol("features")
.setOutputCol("normFeatures")
.setP(2.0)
.transform(assembler1)
normalizer.show()
%spark2
val targetNormalizer = new Normalizer()
.setInputCol("features")
.setOutputCol("normFeatures")
.setP(2.0)
.transform(assembler2)
targetNormalizer.show()Train and evaluate Model
%spark2 val Array(trainingData, testData) = normalizer.randomSplit(Array(0.7, 0.3))
%spark2
val lr = new LinearRegression()
.setLabelCol("bmq_index")
.setFeaturesCol("normFeatures")
.setMaxIter(10)
.setRegParam(1.0)
.setElasticNetParam(1.0)
val lrModel = lr.fit(trainingData)
lrModel.transform(testData).select("features","normFeatures", "bmq_index", "prediction").show() %spark2
println(s"Coefficients: ${lrModel.coefficients} Intercept: ${lrModel.intercept}") %spark2
val trainingSummary = lrModel.summary
println(s"numIterations: ${trainingSummary.totalIterations}")
println(s"objectiveHistory: [${trainingSummary.objectiveHistory.mkString(",")}]")
trainingSummary.residuals.show()
println(s"RMSE: ${trainingSummary.rootMeanSquaredError}")
println(s"r2: ${trainingSummary.r2}") %spark2
val targetTable = lrModel.transform(targetNormalizer).select("date", "estimated_intensity_index", "estimated_fatigue_index", "prediction")
targetTable.show()
Section 4: Save the results in a table
Finally, we will take the results of the prediction and put them in a table called BMQ_PREDICTIONS for later use:
Rename the dataframe to match target columns name
%spark2
val newNames = Seq("date", "estimated_intensity_index", "estimated_fatigue_index", "predicted_bmq")
val targetTableRenamed = targetTable.toDF(newNames: _*)
Delete target table if exists
%jdbc(mysql) drop table if exists BMQ_PREDICTIONS
Write data
%spark2
val prop = new java.util.Properties
prop.setProperty("driver", "com.mysql.jdbc.Driver")
prop.setProperty("user", "bmq_user")
prop.setProperty("password", "Be@stM0de")
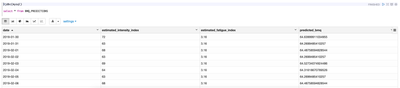
targetTableRenamed.write.mode("append").jdbc("jdbc:mysql://localhost:3306/beast_mode_db", "BMQ_PREDICTIONS", prop)View data
%jdbc(mysql) select * from BMQ_PREDICTIONS