Community Articles
- Cloudera Community
- Support
- Community Articles
- Customizing Atlas (Part2): Deep source metadata & ...
- Subscribe to RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
- Subscribe to RSS Feed
- Mark as New
- Mark as Read
- Bookmark
- Subscribe
- Printer Friendly Page
- Report Inappropriate Content
Created on 12-24-2018 02:57 PM - edited 08-17-2019 05:07 AM
Introduction
In the previous post Customizing Atlas (Part1): Model governance, traceability and registry we:
- provided a brief overview of Atlas types and entities
- showed how to customize Atlas types and entities to fit your own needs and appear in Atlas search and lineage
- customized a special type called model, which inherited from Process and empowered Atlas to govern the deployment of data science models
- commented on operationalizing custom entities
In this post we will:
- customize an Atlas type to represent deep source metadata (metadata beyond source data itself)
- customize an Atlas type to represent devices (metadata about the actual device that generates data)
- embed the device entity in the deep source entity (make the device entity an attribute value in the deep source metadata)
- show how device as an attribute value is a clickable link in the Atlas UI that opens to the full device entity
Concepts and Example
Main Concepts
The main concepts are summarized in the table below.
Example: Gene sequence data ingest
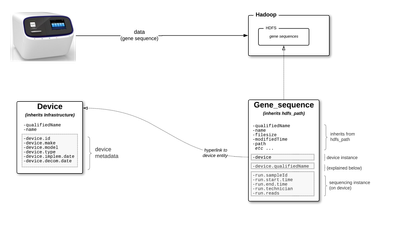
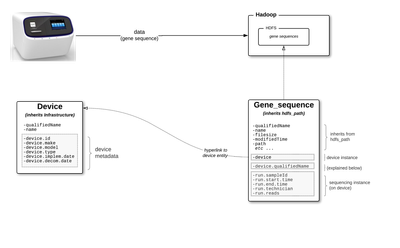
We will use the following scenario to represent these ideas.
- genomics company has multiple gene sequencing devices
- a technician conducts a run on the device, which outputs a blood sample's gene sequence which in turn is ingested to HDFS
- in Atlas, metadata for each device is instantiated as a device entity (Device type inherits from Infrastructure type)
- in Atlas, metadata for each gene sequence is instantiated as a gene_sequence entity (Gene_sequence type inherits from hdfs_path)
- each gene_sequence entity holds deep source metadata (in addition to metadata about the file on hdfs, also metadata about the device the generated the sequence and the specific run on the device, eg the technician's name)
- gene_sequence has an metadata attribute called device, which holds the actual device entity (not string)
Implementation
Device type is implemented as follows (one-time operation):
ATLAS_UU_PWD=$1
ATLAS_HOST=$2
curl -u ${ATLAS_UU_PWD} -ik -H "Content-Type: application/json" -X POST http://${ATLAS_HOST}:21000/api/atlas/v2/types/typedefs -d '{
"enumDefs": [],
"structDefs": [],
"classificationDefs": [],
"entityDefs": [
{
"superTypes": ["Infrastructure"],
"name": "device",
"typeVersion": "1.0",
"attributeDefs": [
{
"name": "qualifiedName",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": true,
"isOptional": false,
"isIndexable": true
},
{
"name": "owner",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "name",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": true,
"isOptional": false,
"isIndexable": true
},
{
"name": "deviceId",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "deviceType",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "deviceModel",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "deviceMake",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "deviceImplemDate",
"typeName": "date",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "deviceDecomDate",
"typeName": "date",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": true,
"isIndexable": true
}
]
}
]
}'
Device entity is instantiated as follows:
ATLAS_UU_PWD=$1
ATLAS_HOST=$2
curl -u ${ATLAS_UU_PWD} -ik -H "Content-Type: application/json" -X POST http://${ATLAS_HOST}:21000/api/atlas/v2/entity/bulk -d '{
"entities": [
{
"typeName": "device",
"attributes": {
"qualifiedName": "Illumina-iSeq100-1092454@gene_sequencer",
"owner": "infra-group",
"name": "Illumina-iSeq100-1092454",
"deviceId": "1092454",
"deviceType": "gene_sequencer",
"deviceModel": "iSeq100",
"deviceMake": "Illumina",
"deviceImplemDate": "2018-08-21T19:49:24.000Z",
"deviceDecomDate": ""
}
}
]
}'
Gene_sequence type is implemented as follows (one-time operation)
ATLAS_UU_PWD=$1
ATLAS_HOST=$2
curl -u ${ATLAS_UU_PWD} -ik -H "Content-Type: application/json" -X POST http://${ATLAS_HOST}:21000/api/atlas/v2/types/typedefs -d '{
"enumDefs": [],
"structDefs": [],
"classificationDefs": [],
"entityDefs": [
{
"superTypes": ["hdfs_path"],
"name": "gene_sequence",
"typeVersion": "1.0",
"attributeDefs": [
{
"name": "qualifiedName",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": true,
"isOptional": false,
"isIndexable": true
},
{
"name": "owner",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "name",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": true,
"isOptional": false,
"isIndexable": true
},
{
"name": "device",
"typeName": "device",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "deviceQualifiedName",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "runSampleId",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "runReads",
"typeName": "int",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "runStartTime",
"typeName": "date",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "runEndTime",
"typeName": "date",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
},
{
"name": "runTechnician",
"typeName": "string",
"cardinality": "SINGLE",
"isUnique": false,
"isOptional": false,
"isIndexable": true
}
]
}
]
}'Gene_sequence entity is instantiated as follows:
ATLAS_UU_PWD=$1
ATLAS_HOST=$2
curl -u ${ATLAS_UU_PWD} -ik -H "Content-Type: application/json" -X POST http://${ATLAS_HOST}:21000/api/atlas/v2/entity/bulk -d '{
"entities": [
{
"typeName": "gene_sequence",
"attributes": {
"clusterName": "prod.genomicsanalytics.com",
"isFile": "true",
"fileSize": "2793872046",
"createdBy": "nifi",
"createTime": "2018-11-12T15:10:03.235Z",
"qualifiedName": "/data/sequence-pipeline/device-output/AB12357@prod.genomicsanalytics.com",
"owner": "jobscheduler",
"name": "/data/sequence-pipeline/device-output/AB12357",
"path": "hdfs://data/sequence-pipeline/device-output/AB12357",
"device": {"uniqueAttributes": {"qualifiedName": "Illumina-iSeq100-1092454@gene_sequencer"}, "typeName": "device"},
"deviceQualifiedName": "Illumina-iSeq100-1092454@gene_sequencer",
"runSampleId": "AB12357",
"runReads": "9",
"runTechnician": "Neeraj Gupta",
"runStartTime": "2018-11-12T09:54:12.432Z",
"runEndTime": "2018-11-12T15:09:59.351Z"
}
}
]
}Results in Atlas UI
Search for gene_sequence entities
We can now search the gene_sequence type and see results (only one result in this example .. and I used the 'Columns' dropdown to customize the result columns).
Notice the Device as a hyperlink.
Drill down to all metadata of a single entity
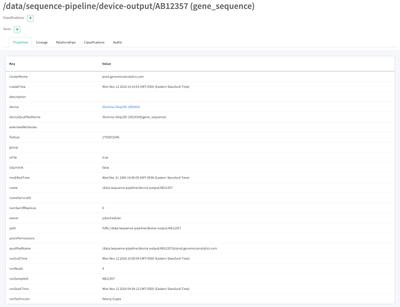
Let's first click the Name to get full list of metadata both inherited from hdfs_path and customized for gene_sequence.
Note that we see the standard hdfs_path properties (like fileSize, path, etc ... for ease of development here, I did not fill in all the values.. this would be done on ingest to the cluster). We also see the device metadata and the run metadata.
Drill down to device metadata
If we click on the link to 'device' (from either of the two places in screenshots above) we see the following.
A note on search and embedded entities
You'll notice that our embedded customized entity 'device' does not show up in search: we cannot directly search by the attributes of an embedded entity (though we can search by on our customized attributes like RunStartTime that uses native Atlas types).
This is the reason I have used the device qualified name (String) as an attribute
Notice how it is constructed: <make>-<model>-<id>@<type>. This allows us to use the search construct 'contains' to search for all gene_sequence entities (ie data in hdfs) that match a device's make, model, id or type, or combination of these.
Summary: What have we accomplished?
This example
For the example here, we know deep knowledge of any gene sequence landed into HDFS. We know:
- gene sequence HDFS path, file size, ingest time etc.
- which device generated the gene sequence data back in the lab
- details of the sample run that was sequenced on the device: technician's name, sample id, how long it took to run the sample, etc
Generalizability
The ideas here can be generalized for you to
- capture any source metadata that goes deeper than the data itself, and
- embed any custom entity as a clickable attribute value in another entity
Use your imagination .. or rather, govern your data deeply.
References
- Atlas Overview
- Using Apache Atlas on HDP3.0
- Apache Atlas
- Altas Type System
- Atlas Rest API
- Related Previous Posts
Acknowledgements
Appreciation to @eorgad and @Former Member rongali for awesome collaboration in the 2018 NE Hackathon which generated many ideas, including those in this post (and for taking first place!).